Natural Products Genomics
Ecological function and application of bacterial natural products
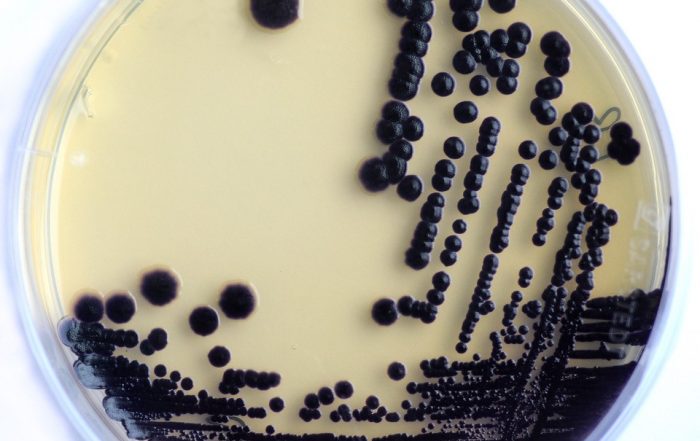
Foto: Dr. Carsten Kegler
Institution: Goethe University Frankfurt
TBG Group members
- Prof. Dr. Helge Bode
- Jürgen Breitenbach, Postdoc
- Edna Bode, Postdoc (affiliated)
- Carsten Kegler, Postdoc (affiliated)
- Kenan Bozhüyük, Postdoc (affiliated)
- Peter Grün, TA
- Timo Obermeier, PhD student (affiliated)
- Leonard Präve, PhD student (affiliated)
SUMMARY
Several clinically used drugs like antibiotics or anti-cancer and immune-suppressive drugs are produced by bacteria as natural products or specialized metabolites (SMs). While these SMs are important for our health care system, we hardly know why the bacteria produce them or what other bacterial SMs await their identification and future clinical application. The focus of our project is to develop molecular tools to identify novel SMs faster, manipulate the underlying biosynthetic gene clusters (BGCs) in order to produce novel derivatives of these SMs, and to elucidate regulatory mechanisms that influence SM production in bacteria. Especially with respect to the translational aspect of this reseach, the work with bacterial genomes is essential since only they offer the possibility of a manipulation in the desired detail.
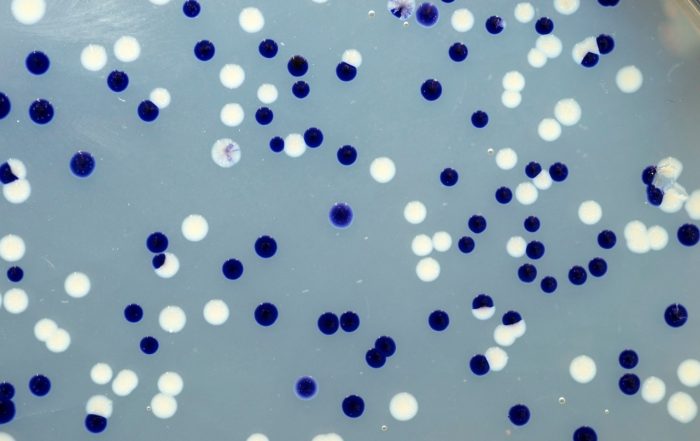
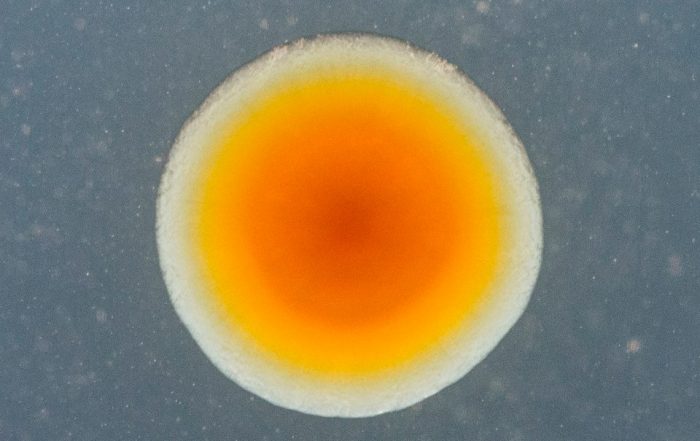
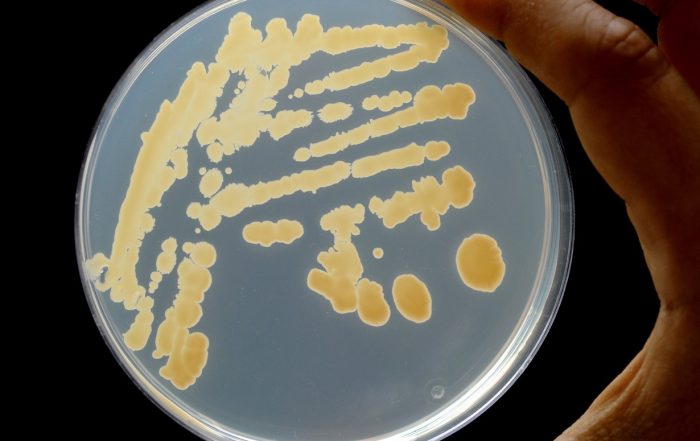
Fotos: Dr. Carsten Kegler
GROUP EXPERTISE / METHODS
- Identification and structure elucidation of natural products
- manipulation of bacterial genomes
- mass spectrometry
- biochemistry
- molecular microbiology
ORGANISMS / GENOMES ANALYZED
- Photorhabdus
- Xenorhabdus
- Pseudomonas
- nematodes
- insect-associated bacteria
PUBLICATIONS
Machado, R.A.R., Thönen, L., Arce, C.C.M. et al. Engineering bacterial symbionts of nematodes improves their biocontrol potential to counter the western corn rootworm. Nat Biotechnol 38, 600–608 (2020). https://doi.org/10.1038/s41587-020-0419-1
Bode E, Heinrich AK, Hirschmann M, Abebew D, Shi YN, Vo TD, Wesche F, Shi YM, Grün P, Simonyi S, Keller N, Engel Y, Wenski S, Bennet R, Beyer S, Bischoff I, Buaya A, Brandt S, Cakmak I, Çimen H, Eckstein S, Frank D, Fürst R, Gand M, Geisslinger G, Hazir S, Henke M, Heermann R, Lecaudey V, Schäfer W, Schiffmann S, Schüffler A, Schwenk R, Skaljac M, Thines M, Thines E, Ulshöfer T, Vilcinskas A, Wichelhaus TA, Bode HB, Promoter activation in hfq mutants as an efficient tool for specialized metabolite production enabling direct bioactivity testing. Angew Chem Int Ed 2019, 58 (52), 18957-18963. VIP paper. https://doi.org/10.1002/anie.201910563
Berry, D., Mace, W., Grage, K., Wesche, F., Gore, S., Schardl, C., Young, C., Dijkwel, P., Leuchtmann, A., Bode, H., and Scott, B. Efficient nonenzymatic cyclization and domain shuffling drive pyrrolopyrazine diversity from truncated variants of a fungal NRPS. PNAS 116 (51) 25614-25623 (2019). https://doi.org/10.1073/pnas.1913080116
Wang, G., Zhao, Z., Ke, J. et al. CRAGE enables rapid activation of biosynthetic gene clusters in undomesticated bacteria. Nat Microbiol 4, 2498–2510 (2019). https://doi.org/10.1038/s41564-019-0573-8
Schmidt, R., Ulanova, D., Wick, L.Y. et al. Microbe-driven chemical ecology: past, present and future. ISME J 13, 2656–2663 (2019). https://doi.org/10.1038/s41396-019-0469-x
Shi, Y., Pusch, S., Shi, Y., Richter, C., Maciá-Vicente, J., Schwalbe, H., Kaiser, M., Opatz, T., and Bode, H. (±)-Alternarlactones A and B, Two Antiparasitic Alternariol-like Dimers from the Fungus Alternaria alternata P1210 Isolated from the Halophyte Salicornia sp., The Journal of Organic Chemistry 2019 84 (17), 11203-11209. https://doi.org/10.1021/acs.joc.9b01229
Bozhüyük KAJ, Linck A, Tietze A, Kranz J, Wesche F, Nowak S, Fleischhacker F, Shi YN, Grün P, Bode HB, Modification and de novo design of non-ribosomal peptide synthetases using specific assembly points within condensation domains. Nat Chem. 2019 11(7): 653-661. https://doi.org/10.1038/s41557-019-0276-z