Genomic Biomonitoring
eDNA Chips
Institution: Conservation Genetics Section, Senckenberg Society for Nature Research (Gelnhausen station)
Main task of the project “eDNA chip” is the development of several microfluidic arrays for rapid and cost-effective species detection based on environmental samples with low DNA content, such as water, scats, or hairs. In a first step, we develop two detection chips for native fish and amphibian species, which will be applied in several applied monitoring studies and compared with traditional surveys. Additional SNP chips are in preparation for individual discrimination and population monitoring of several endangered species of the habitats directive, such as European bison and hazel dormouse. Our ultimate goal is to transfer these reduced genomic detection platform into monitoring and conservation practice.
- Genotyping of forensic and environmental samples based on microsatellites and SNPs
- Microfluidic SNP genotyping (Fluidigm)
- molecular population genetics
- conservation genetics/genomics
- wildlife biology
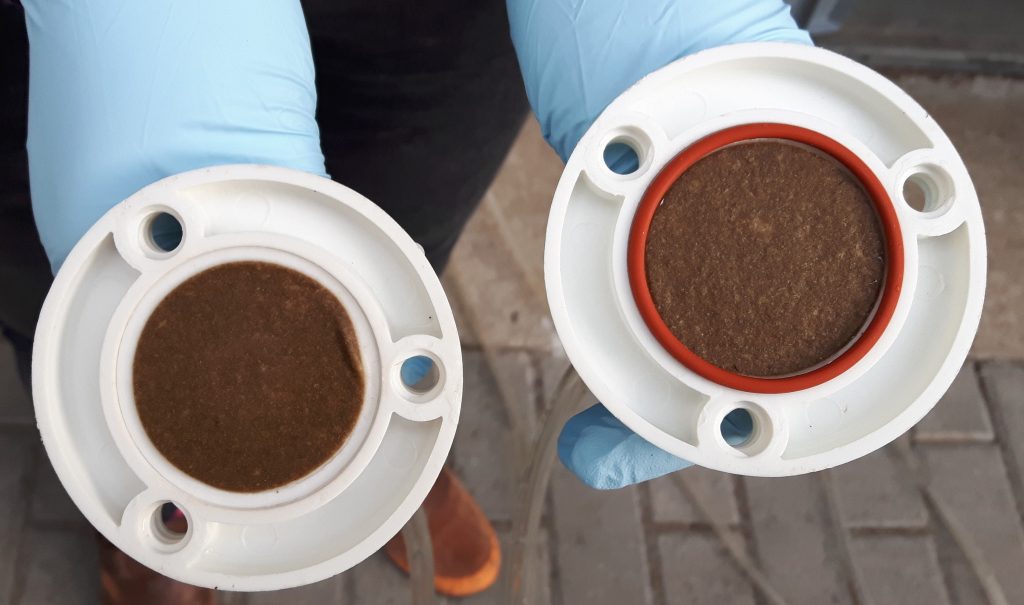






Water is sampled via filters with different pore sizes, here using a glass fiber filter with a pore size of 2 µm. Once the maximum filtration capacity is reached, the filters are removed from the filter housing and are ready for subsequent eDNA purification and molecular biology analyses.
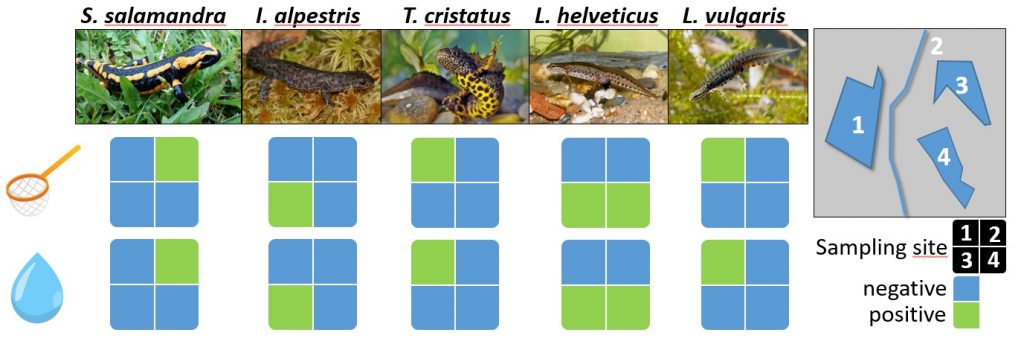
Detection of native newts: classic monitoring vs. eDNA chip in four ponds
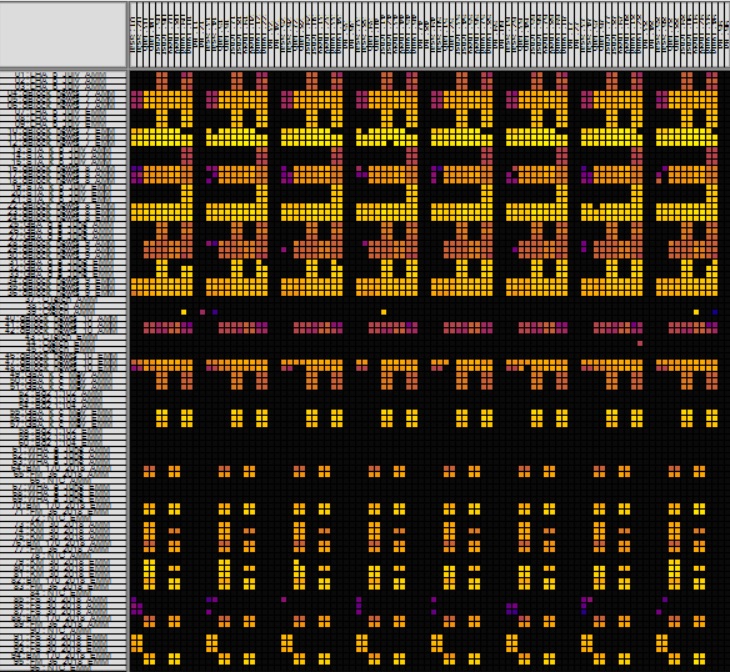
Heatmap view of raw data obtained from a run with the Biomark HD microfluidic system (FluidigmTM) of eDNA samples of native newt species.
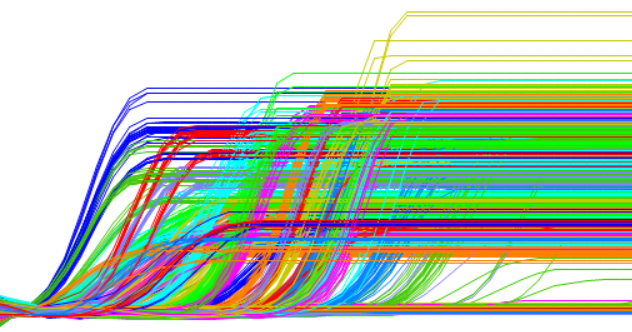

Microfluid technology allows the simultaneous analysis of thousands of real-time PCR reactions within a single chip. This saves time and expenses and minimizes the consumption of sample material.
PUBLICATIONS
von Thaden A, Nowak C, Tiesmeyer A, et al. (2020). Applying genomic data in wildlife monitoring: Development guidelines for genotyping degraded samples with reduced single nucleotide polymorphism panels. Molecular Ecology Resources 20(3), 662-680.
Harmoinen J, von Thaden A, Aspi J, et al. (2021). Reliable wolf-dog hybrid detection in Europe using a reduced SNP panel developed for non-invasively collected samples. BMC Genomics 22, 473.